Broughton 2015 Abstract MiPschool Greenville 2015
| Natural selection on OXPHOS proteins varies among organisms with differences in aerobic performance. |
Link:
Broughton RE Zhang F (2015)
Event: MiPschool Greenville 2015
Oxidative phosphorylation (OXPHOS) is the primary source of ATP in eukaryotes and serves as a mechanistic link between variation in genotypes and energetic phenotypes. Increased aerobic capacity may be facilitated by numerous factors, including increased expression of OXPHOS genes, increase in number and morphology of mitochondria, enhanced O2 transport within the body, and structural variation in OXPHOS proteins. Directional (positive) natural selection on particular amino acid sites may reveal important functional variation in OXPHOS proteins. Although evidence of natural selection on mitochondrial OXPHOS genes has been documented in several animal groups, the relationship between selection and OXPHOS function has not been well characterized. Because organismal performance should be a function of OXPHOS functional efficiency, we investigated this relationship by examining patterns of natural selection on OXPHOS genes in several vertebrate lineages with extreme differences in aerobic performance.
We assembled OXPHOS genes for several groups of taxa, including 40 mammal species from diverse taxonomic orders as well as trees for several fish groups each including one or more orders. For the mammals, all OXPHOS genes (both mitochondrial and nuclear encoded) were examined and species represented a range of energetic habits. In fishes, mitochondrial and some nuclear genes were examined from largely sedentary groups as well as highly active, high-performance swimmers. We used estimates of the synonymous (dS) to non-synonymous (dN) amino acid substitution rate ratios to provide evidence of natural selection. Directional (positive) selection was inferred where the dN/dS ratio was greater than 1.0, while purifying (negative) selection was indicated by dN/dS ratios much less than 1.0. Specific amino acid sites under positive selection were identified using a Bayesian procedure and for several groups these were localized on the 3-dimensional structure of their respective OXPHOS complexes.
We found that patterns of natural selection on OXPHOS genes are complex among fishes and mammals with different locomotive performance or other energetic demands. Positive selection is associated with several high performance groups but is also associated with some low performance taxa. For example among mammals positive selection was detected for many genes on the horse and primate lineages but was also detected on the lineage leading to sloths and armadillos. Some high performance taxa such as dolphins did not exhibit evidence of positive selection. In fishes positive selection was strongest on low-performance seahorses and flounders, while the high-performance tunas and marlins exhibited strong purifying (negative) selection. Thus, positive natural selection was not consistently associated with high performance taxa and in several cases selection was quite strong on low-performance taxa. Within some high-performance lineages, negative selection in the form of functional constraint appears to predominate.
Labels: MiParea: Respiration, mtDNA;mt-genetics
Organism: Horse, Other mammals, Fishes
Event: Poster
Affiliation
Oklahoma Biol Survey and Dept Biol, Univ Oklahoma, Norman, OK, USA. - [email protected]
Abstract continued
We provide evidence that selection on OXPHOS protein subunits varies in both form and intensity within and among lineages of vertebrates. The type and direction of selection vary in ways that would not be readily predicted based solely on organismal performance. These results provide empirical evidence for fluctuating selection on OXPHOS associated with divergence in aerobic performance. The broader pattern indicates a complex interplay between organismal adaptations, ATP demand and OXPHOS function and underscores the need to understand genetic variation among taxa in relation to specific OXPHOS functional performance.
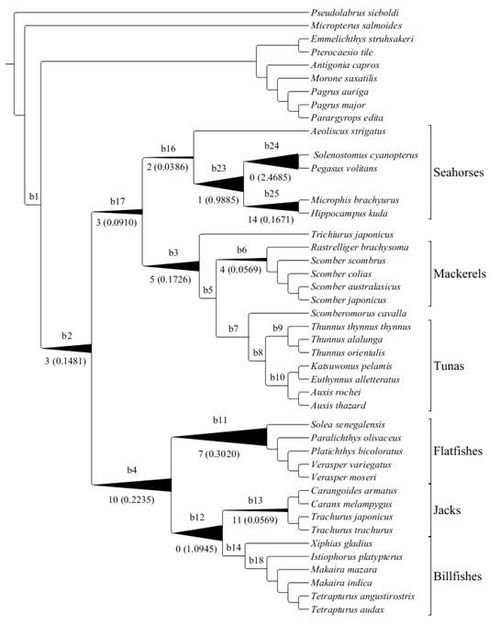
Figure 1
Figure 1. Positive selection analysis on mitochondrial genes on each branch (arbitrary labels, b1-b24, appear above each branch). The number below each branch represents the number of positively selected sites with posterior probability higher than 0.8. The number in parentheses is the product of the dN/dS ratio and the proportion of sites with a ratio >1. These values serve as indicators of the strength of positive selection. The width of the branches is proportional to the strength of positive selection (because ω2 for b12 and b24 is extremely high, the width was capped for ease of visualization). Labled branches without values listed have no evidence of positive selection.