Koopman 2017 Abstract MITOEAGLE Barcelona
| Standardized quantification of mitochondrial morphology and function in living patient fibroblasts using microscopy. Koopman_Presentation |
Link: MitoEAGLE
Koopman Werner JH (2017)
Event: MitoEAGLE Barcelona 2017
Mitochondria play a central role in cellular (patho)physiology and display a highly variable morphology that is probably coupled to their functional state [1]. Our research aims to quantitatively understand the molecular connection between mitochondrial metabolism and (ultra)structure with particular attention to redox signaling and biomolecule diffusion. To this end we study various cell models and a mouse model of mitochondrial complex I (CI) deficiency. As a key technology, protein-based and chemical fluorescent reporter molecules are introduced in the cells and their signals are quantified using life cell microscopy, image processing/quantification and data mining. Protein diffusion is studied by combining photobleaching strategies, single-molecule spectroscopy and in silico techniques. In primary fibroblasts from Leigh Syndrome (LS) patients, isolated CI deficiency is associated with mitochondrial morphological and functional changes and increased reactive oxygen species (ROS) levels. Here we present a protocol [2] allowing unbiased and automated quantification of mitochondrial “morphofunction” (i.e. morphology and membrane potential), cellular parameters (size, confluence) and nuclear parameters (number, morphology) in intact living Primary Human Skin Fibroblasts (PHSFs). Cells are cultured in 96-well plates and stained with tetramethyl rhodamine methyl ester (TMRM), Calcein-AM and Hoechst 33258 (Fig. 1). Next, multispectral fluorescence images are acquired using automated microscopy and processed to extract 44 descriptors. Subsequently, the descriptor data is subjected to a quality control (QC) algorithm based upon Principal Component Analysis (PCA) and interpreted using univariate, bivariate and multivariate analysis. The protocol requires a time investment of about 4 hours distributed over 2 days. Although specifically developed for PHSFs that are widely used in preclinical research, the protocol is portable to other cell types and can be up-scaled for implementation in High-Content Screening (HCS). Using a machine learning strategy [3], we discriminated between fibroblasts of a healthy individual and an LS patient based upon their mitochondrial morpho-functional phenotype. This allowed evaluating the therapeutic potential of newly developed cellular intervention strategies [4].
• Bioblast editor: Kandolf G
• O2k-Network Lab: NL Nijmegen Koopman WJ
Labels: Pathology: Inherited
Organism: Human Tissue;cell: Fibroblast Preparation: Intact cells Enzyme: Complex I
Event: A1
Affiliations
- Dept Biochemistry, Live-cell Bioenergetics subgroup, Radboudumc, Nijmegen, The Netherlands.- [email protected]
Figure 1
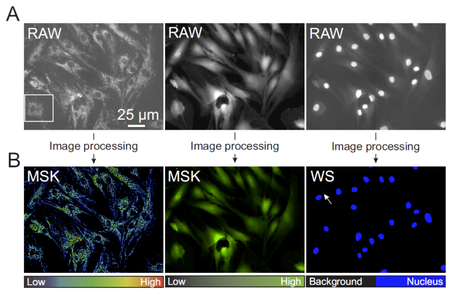
Figure 1. Overall image quantification strategy. (A) Typical example of a TMRM, Calcein and Hoechst image (from left to right) obtained from the same well (RAW images). (B) Images obtained after image processing (MSK=masked image, WS=after watershed operation).
References
- Willems PHGM, Rossignol R, Dieteren CEJ, Murphy MP, Koopman WJH (2015) Redox homeostasis and mitochondrial dynamics. Cell Metab 22:207-18.
- Iannetti EF, Smeitink JAM, Beyrath J, Willems PHGM, Koopman WJH (2016) Multiplexed high-content analysis of mitochondrial morphofunction using live-cell microscopy. Nature Protoc 11:1693-1710.
- Blanchet L, Smeitink JAM, Breuer ME, van Emst - de Vries SE, Vogels C, Pellegrini M, Jonckheere AI, Rodenburg RJT, Buydens LMC, Beyrath J, Willems PHGM, Koopman WJH (2015) Analysis of small molecule phenotypic effects using combined mitochondrial morpho-functional fingerprinting and machine learning. Sci Rep 5:8035.
- Koopman WJH, Beyrath J, Fung W, Koene S, Rodenburg RR, Willems PHGM, Smeitink JAM (2016) Mitochondrial disorders in children: toward development of small-molecule treatment strategies. EMBO Mol Med 8:311-27.