Takaharu 2015 Abstract MiP2015
| Higd1a is a positive regulator of cytochrome c oxidase. |
Link:
Takaharu H, Asano Y, Shintani Y, Kioka H, Tsukihara T, Yoshikawa S, Takashima S (2015)
Event: MiP2015
Cytochrome c oxidase (CcO) is the only enzyme that utilizes oxygen to produce a proton gradient for ATP production in mitochondrial oxidative phosphorylation. Mammalian CcO is composed of 13 different subunits containing four redox active metal centers [1]. Because CcO is the only enzyme in the body that can utilize oxygen for energy transduction, it has been suggested that regulatory mechanism of CcO is dependent on oxygen concentration [2]. In this study, we aimed to identify CcO regulator which induced under hypoxia.
We screened gene expression profiles of neonatal rat cardiomyocytes and found Higd1a as one of an up-regulating genes in hypoxia. Biochemical analysis revealed that Higd1a directly binds to CcO and structural analysis by resonance Raman revealed that Higd1a caused structural changes in the CcO, especially around heme a, the active center that drives proton-pump [3]. Endogenous induction of Higd1a in rat cardiomyocytes under hypoxia increased mitochondrial ATP synthesis. Moreover, exogenous Higd1a successfully improved cell survival in rat cardiomyocytes.
By an identification of Higd1a and its biochemical and structural investigation, we demonstrated that Higd1a could exhibit an increase of ATP production via interaction with CcO.
Labels: MiParea: mtDNA;mt-genetics
Stress:Oxidative stress;RONS Organism: Rat Tissue;cell: Heart
Enzyme: Complex IV;cytochrome c oxidase
Event: B1, Oral
MiP2015
Affiliations
1-Dept Cardiovascular Med; 2-Dept Med Biochem, Osaka Univ Graduate School Med, Suita, Osaka; 3-Dept Life Sc, Univ Hyogo, Japan. - [email protected]
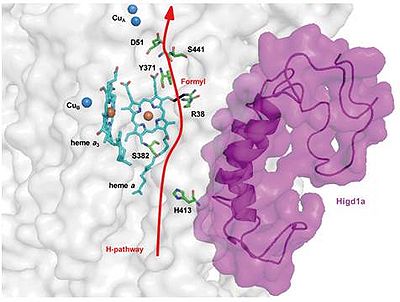
Figure 1
Figure 1. Higd1a acts on the H-pathway. Model depicting our docking simulation (side view) and its relationship with the H-pathway. The model shows the location of Higd1a (magenta) in the CcO complex (white) and its relationship to R38 of cytochrome c oxidase subunit I and the formyl group of heme a, a component of the H-pathway (red arrow).
References and acknowledgements
- Tsukihara T, Aoyama H, Yamashita E, Tomizaki T, Yamaguchi H, Shinzawa-Itoh K, Nakashima R, Yaono R, Yoshikawa S (1996) The whole structure of the 13-subunit oxidized cytochrome c oxidase at 2.8 A. Science 272:1136-44.
- Vukotic M, Oeljeklaus S, Wiese S, Vogtle FN, Meisinger C, Meyer HE, Zieseniss A, Katschinski DM, Jans DC, Jakobs S, Warscheid B, Rehling P, Deckers M (2012) Rcf1 mediates cytochrome oxidase assembly and respirasome formation, revealing heterogeneity of the enzyme complex. Cell metabolism 15:336-47.
- Muramoto K, Ohta K, Shinzawa-Itoh K, Kanda K, Taniguchi M, Nabekura H, Yamashita E, Tsukihara T, Yoshikawa S (2010) Bovine cytochrome c oxidase structures enable O2 reduction with minimization of reactive oxygens and provide a proton-pumping gate. Proceedings Nat Acad Sc USA 107:7740-5.