Template:SUIT-024 O2 ce-pce D056: Difference between revisions
From Bioblast
Garcia Luiz (talk | contribs) (Created page with "right|190px|link=http://www.bioblast.at/index.php/MitoPedia:_SUIT |MitoPedia: SUIT == Steps and respiratory states == File:Ce1;1Dig;1PM;2T;2D;3Omy...") |
Beno Marija (talk | contribs) No edit summary |
||
| (7 intermediate revisions by 2 users not shown) | |||
| Line 1: | Line 1: | ||
[[Image:SUIT-MitoFit.png|right|190px|link=http://www.bioblast.at/index.php/MitoPedia:_SUIT |MitoPedia: SUIT]] | [[Image:SUIT-MitoFit.png|right|190px|link=http://www.bioblast.at/index.php/MitoPedia:_SUIT |MitoPedia: SUIT]] | ||
== Steps and respiratory states == | == Steps and respiratory states == | ||
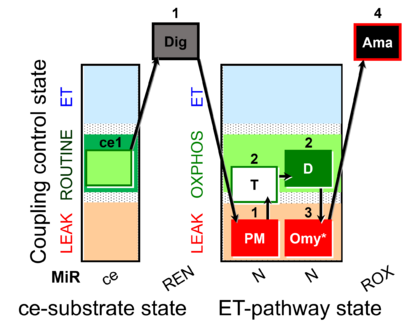
[[File:Ce1;1Dig;1PM;2T;2D;3Omy;4Ama.png| | [[File:Ce1;1Dig;1PM;2T;2D;3Omy;4Ama.png|410px]] | ||
{{Template:SUIT cells}} | {{Template:SUIT cells}} | ||
| Line 20: | Line 20: | ||
|- | |- | ||
| 2T | | 2T | ||
| [[PM]]<sub>''P''</sub> | | [[PM]]<sub>''L''</sub> or [[PM]]<sub>''P''</sub> | ||
| [[N]] | | [[N]] | ||
| CI | | CI | ||
| Line 26: | Line 26: | ||
*{{Template:SUIT N}} | *{{Template:SUIT N}} | ||
* | * In the absence of ATPase activity, the [[LEAK respiration|LEAK]] state is maintained. However, if ATPases are active and thus generate ADP, respiration coupled to phosphorylation is stimulated. Stimulation is limited up to [[OXPHOS]] capacity; therefore, higher ATPase activities cannot be determined. | ||
|- | |- | ||
| Line 40: | Line 40: | ||
|- | |- | ||
| 3Omy | | 3Omy | ||
| | | [[PM]]<sub>''L''(Omy)</sub> | ||
| | | | ||
| | | | ||
| 1PM;2T;2D;3Omy | | 1PM;2T;2D;3Omy | ||
*{{Template:SUIT L(Omy)}} | *{{Template:SUIT L(Omy)}} | ||
|- | |- | ||
| Line 57: | Line 56: | ||
|} | |} | ||
{{Keywords: SUIT protocols}} | |||
{{ | |||
Latest revision as of 12:35, 25 November 2020
Steps and respiratory states
| Step | State | Pathway | Q-junction | Comment - Events (E) and Marks (M) |
|---|---|---|---|---|
| ce1 | ROUTINE | ce1
| ||
| 1Dig | REN | ce1;1Dig
|
| Step | State | Pathway | Q-junction | Comment - Events (E) and Marks (M) |
|---|---|---|---|---|
| 1PM | PML(n) | N | CI | 1PM
|
| 2T | PML or PMP | N | CI | 1PM;2T
|
| 2D | PMP | N | CI | 1PM;2T;2D
|
| 3Omy | PML(Omy) | 1PM;2T;2D;3Omy
| ||
| 4Ama | ROX | 1PM;2T;2D;3Omy;4Ama
|
- Bioblast links: SUIT protocols - >>>>>>> - Click on [Expand] or [Collapse] - >>>>>>>
- Coupling control
- Pathway control
- Main fuel substrates
- » Glutamate, G
- » Glycerophosphate, Gp
- » Malate, M
- » Octanoylcarnitine, Oct
- » Pyruvate, P
- » Succinate, S
- Main fuel substrates
- Glossary